Mobile Health systems
Mobile Ultrasonic Sonar Exercise Sensing

Physical inactivity is the fourth leading risk factor for death worldwide, and yet eighty percent of US adults do not meet national exercise recommendations. In collaboration with the Sports Institute at UW Medicine, the UbiComp Lab is designing a smartphone application that can be prescribed to patients at University of Washington Medicine clinics. Physical activity is a vital sign that should be monitored regularly alongside blood pressure and BMI. Our application is designed to assist in goal setting, provide context-aware nudges, and connect exercise data with health care providers. To extend the quantification of physical activity beyond step counting, I evaluated the use of acoustic sonar sensing to classify a wide range of exercises. I mentored two high school interns who are assisted with data collection and analysis.
Ubiquitous Non-Invasive Osteoporosis Screening

Osteoporosis is characterized by a decrease in bone mass density (BMD) causing millions of fractures annually. The clinical standard for measuring BMD requires radiation, expensive equipment, and trained staff, motivating the need for a low-cost and ubiquitous alternative. I prototyped OsteoApp, a smartphone application that aims to infer bone mass density by measuring resonant frequency in a human tibia using the built-in accelerometer. I tested this system alongside a low-noice accelerometer data collection system I built with patients from local retirement communities as well as a control group of healthy volunteers, and performed signal processing and data analysis in Python. I presented this work at the 2019 University of Washington Undergraduate Research Symposium.
PosterReal-time Smartphone Pulse Transit Time Measurement
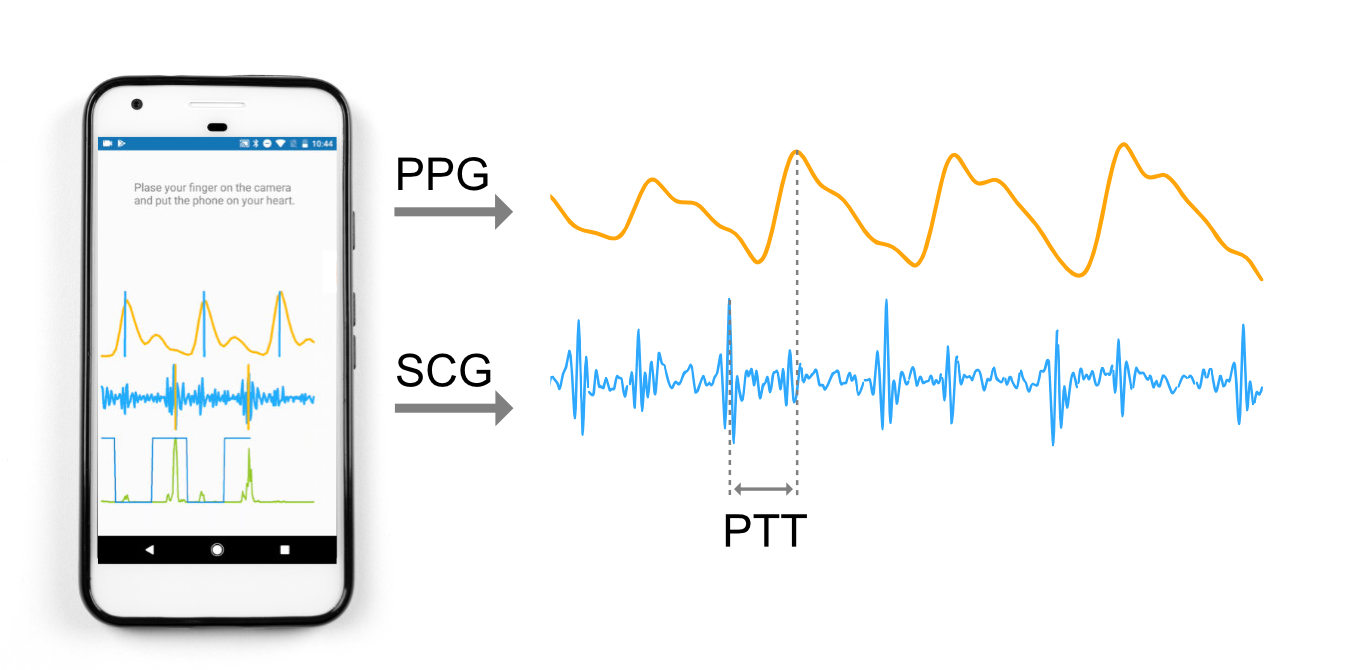
Pulse transit time (PTT) is the time differential between the heartbeat and the arrival of the pulse pressure wave at the fingertip. Since PTT is indicative of arterial stiffness, it can provide an early sign of atheroscloerosis and cardiovascular disease. Additionally, PTT is inversely correlated with blood pressure, providing an opportunity to perform noninvasive estimation of of blood pressure using commodity devices. Prior work from my lab proposed the use of smartphones to measure PTT; I built upon this work by implementing a smartphone application that can perform real-time sensing, signal processing, and visualization code for the Android smartphone platform. I presented this work within the Paul G. Allen School of Computer Science & Engineering at an annual technology CEO summit and the 2018 Industry Affiliates Research Day, as well as the 2018 University of Washington Undergraduate Research Symposium.
Wearable Physiological Sensing
Earbud Physiological Sensing

With wireless earbuds becoming increasingly powerful and widely used, we recognize an opportunity to leverage them as a platform for performing continuous physiological measurements. Modern earbuds can contain acoustic, inertial, optical, and proximity sensors that may be re-purposed to track vital signs and perform diagnostics. My contribution to this work includes prototyping signal processing techniques for in-ear physiological sensing, in addition to supporting study design, data collection, data analysis, and paper writing.
Multi-Channel Facial Photoplethysmography

The face provides a very unique opportunity for performing physiological sensing using wearables and camera-based systems. Measuring the vascular network in the face may enable disease diagnosis and continuous monitoring. A powerful and widely used technique for facial sensing is photoplethysmography (PPG), a non-invasive optical measurement of blood pulses. However, facial PPG remains under-explored in terms of both signal acquisition and analysis. To characterize the timing and morphological features of facial PPG waveforms I designed, built, and tested a multi-channel facial PPG sensing system that can record synchronized pulse waveforms at multiple locations and optical wavelengths. I worked closely with a clinical collaborator from University of Washington Medicine, in addition to mentoring two undergraduate students.
PaperPopulation Health and Assay Automation
SARS-CoV-2 Screening in Public Transportation Air Filtration
Motivated by the need for population level monitoring of COVID-19 transmission, the UbiComp Lab partnered with Microsoft Research to develop protocols for obtaining environmental samples from public transportation air filtration systems to detect the presence of SARS-CoV-2. I supported these efforts by advising on study design, performing literature review, coordinating with collaborators, and participating in paper writing.
PaperStreamlining SARS-CoV-2 Molecular Assays
Detecting and mitigating outbreaks of COVID-19 requires rapid and high throughput testing, disproportionately impacting regions with limited access to reagents, supplies, and trained staff. I worked with bioengineering collaborators to develop and test faster and simpler COVID-19 protocols by performing direct amplification, bypassing the RNA extraction step. I created image processing algorithms for quantifying output fluorescence to reduce time and human error for point-of-care COVID-19 testing applications. This system was deployed for testing at clinical facilities in the US and Zimbabwe.
PaperComputer Vision for HIV Drug Resistance Testing
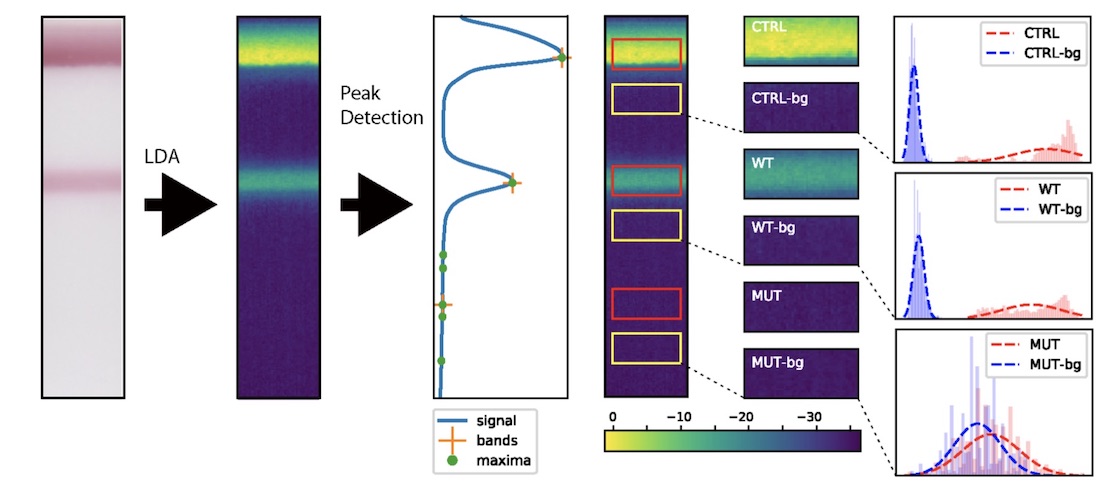
Testing for drug resistant strains of HIV is necessary for clinicians to effectively treat patients; however, the standard genotyping assays like Sanger sequencing are infeasible in resource-limited settings where drug resistant HIV is increasingly circulating. OLA-Simple is a low-cost paper-based lateral flow strip test and chemistry kit that can be used to amplify and detect low amounts of drug-resistant strains of HIV. Five common drug-resistant mutations can be visualized as colored bands on a paper strip; however, human error limits the sensitivity and specificity of this test when interpreted by eye. I built computer vision code to read flatbed scanner images, isolate paper strips, and measure the band intensities to interpret the test results. I used this code to generate data for major tables and figures in the paper for this project, and performed an evaluation with training and testing datasets that demonstrated over 99% accuracy.
Near Point-of-Care Assay for HIV Drug Sensitivity
Abacavir is a nucleoside analog reverse-transcriptase inhibitor used to treat HIV. Although it is generally well tolerated, it triggers dangerous anaphylactic shock in patients with the HLA-B*57:01 genotype. Scientists and engineers from UW Bioengineering and Seattle Children’s Hospital developed a colorimetric lateral flow assay to identify patients at risk for drug sensitivity. I supported this work by applying image processing to reduce human error in assay interpretation.
Paper